Ming-Ko Chiang
MING-KO, CHIANG
Tel: (05)2720411 ext. Office 66505 Lab 61514
Fax: (05)2722871
Email:biomkc@ccu.edu.tw
- 1990-1997 Ph.D. Program of Cell and Developmental Biology, Harvard University, Massachusetts, USA.
- 1982-1986 B.S. Department of Agricultural Chemistry, National Taiwan University, Taipei, Taiwan.
- 2012/8- present Association Professor, Department of Biomedical Sciences, National Chung Cheng University
- 2004/8-2012/7 Assistant Professor, Department of Life Science, National Chung Cheng University
- 2002-2004 Postdoctoral Research Fellow,
- Orthopaedic Research Lab, University of California at Davis, California, USA.
- 1997-2002 Postdoctoral Research Fellow,
- Department of Molecular and Cellular Biology, Harvard University, Massachusetts, USA.
- 1986-1987 Research Assistant, Development Center for Biotechnology, Taipei, Taiwan.
- 1997 Howard Hughes Medical Institute Postdoctoral Fellowship
- 2000 The Medical Foundation Fellowship, MA.
Pancreas Development
My research is focused on the molecular mechanisms that regulate the development of the pancreas. I am particularly interested in the pathways that specify pancreatic cell fate and control pancreatic growth. Defects in these mechanisms may contribute to diseases such as diabetes mellitus and pancreatic cancer. My long-term goal is to identify pancreatic stem cells and other progenitor cells, and to compare the gene expression profiles of these cells with each other and with their differentiated progeny. With this approach, many components of the genetic networks that regulate pancreatic development would be identified, and their specific functions would be further characterized. This research will provide information necessary to control the behavior of the pancreatic cells in vitro and in vivo, and lead to alternative treatments for diabetic patients. In addition, by comparing the gene expression profiles of the pancreatic stem cells and other types of stem cells, we should be able to identify common genetic requirements for cellular self-renewal.
Genetic networks that regulate the pancreatic development
Aim #1
During early embryogenesis, the single layer of midgut endoderm epithelium evaginates to form the pancreatic buds (one dorsal and one ventral). These buds go on to proliferate, branch, and differentiate. The molecular pathways that control these processes are largely unknown. By combining custom DNA microarray analysis with modified single cell PCR, I have developed a system to study the gene expression profiles of single cells at different embryonic stages. With this technique, I have identified pancreatic cells at different stages of differentiation. In addition, these cells can be aligned in a developmental pathway (Fig.1). Among these cells, a particular population of cells that unexpectedly co-express transcription factors PDX-1, Nkx2.2, Nkx6.1, and p48 may represent multi-potent progenitor cells. I plan to extensively characterize the gene expression profiles of these different cell types, and to identify specific signaling molecules or transcription factors required for growth and differentiation of stem cells or progenitor cells within the pancreas.
Aim #2
In addition, I have used the single cell PCR products of different pancreatic cell types for small-scale differential screening. I have identified a numbers of novel genes and confirmed their specific expression in developing pancreas by in situ hybridization (Fig.2). Some of these genes encode molecules similar to known proteins, and some of them are not homologous to any known genes. The specific functions of these genes will be studied in cultured pancreatic cells or in genetically manipulated (transgenic or knock-out) mice.
Fig. 1A-1I. Various Pancreatic Cell types aligned in the pancreas developmental pathway.
The upper panel of each figure shows the hybridization result of the custom microarray with probes made from various pancreatic single cells. The lower panel of each figure summarizes the combination of pancreatic genes expressed in each particular type of cell. Fig. 1A, 1B, 1C, and 1D illustrate the sequence of exocrine cell differentiation during pancreas development. Fig. 1A, 1B, 1E, 1F, 1G, 1H, and 1I illustrate the sequence of endocrine cell differentiation. Cells shown in Fig. 1A and 1B may represent the common progenitor cells for both pancreatic exocrine and endocrine cells.
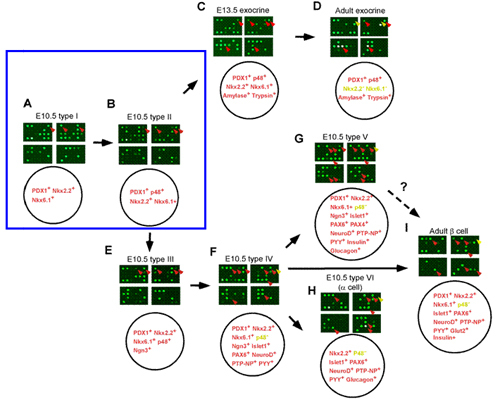
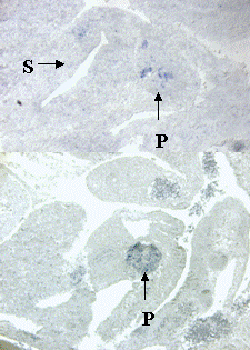
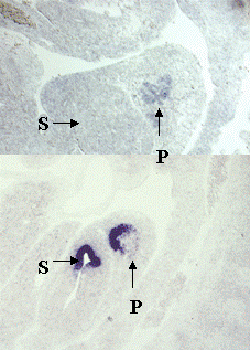
Figure 2. Specific expression patterns of novel genes that were identified by differential screening using single cell PCR products.
A-D, In situ hybridization with probes made from four sample novels genes on sagittal sections of E10.5 embryos. Note that each gene has a specific expression pattern within the pancreas. P, Pancreas; S, Stomach.
Chiang, M.-K. and Flanagan, References J. G. (2002) Nucleic acids encoding neural/pancreatic receptor tyrosine phosphatase. United States Patent No. 6,399,326
- Chiang, M. K., Lu, M. C., Liu, L. C., Lin, C. T. and Lai, Y. C., 2011,Impact of Hfq on Global Gene Expression and Virulence in Klebsiella pneumoniae., PLoS One, Vol.6, No.7, e22248。(SCI)
- Lin, Y. C., Lu, M. C., Tang, H. C., Liu, H. C., Chen, C. H., Liu, K. S., Lin, C., Chiou, C. S., Chiang, M. K., Chen, C. M. and Lai, Y. C.,2011, Assessment of hypermucoviscosity as a virulence factor for experimental Klebsiella pneumoniae infections: comparative virulence analysis with hypermucoviscosity-negative strain, BMC Microbiology, Vol.11, No.50。(SCI)
- Shen, C. H., Wu, J. D., Hsu, C. D., Jou, Y. C., Lin, C. T., Wang, M., Wu, S. F., Chan, M. W., Chiang, M. K., Fang CY, and Chang D. The high incidence of JC virus infection in urothelial carcinoma tissue in Taiwan. Journal of Medical Virology 83:2191-9, 2011
- Tu, Y. C.,Lu M. C., Chiang, M. K., Huang, S. P., Peng, H. L., Chang, H. Y., Jan, M. S. and Lai, Y. C., 2009. Genetic Requirements for Klebsiella pneumoniae-Induced Liver Abscess in an Oral Infection Model. Infect Immun. 77(7):2657-71
- Chiang, M. K., Liao, Y. C., Kuwabara Yasuko, and Lo, S. H. (2005) Inactivation of tensin3 in mice results in growth retardation and postnatal lethality. Developmental Biology 279: 368-377. (SCI, Impact Factor: 5.351)
- Chiang, M. K. and Melton, D. A. (2003) Single cell transcript analysis of pancreas development. Developmental Cell 4: 383-393.
- Bergemann, A. D., Zhang, L., Chiang, M. K., Brambilla, R., Klein R., and Flanagan, J. G. (1998) Ephrin-B3, a ligand for the receptor EphB3, expressed at the midline of the developing neural tube. Oncogene 16: 471-480.
- Chiang, M. K. and Flanagan, J. G. (1996) PTP-NP, a new member of the receptor protein tyrosine phosphatase family, implicated in development of nervous system and pancreatic endocrine cells. Development 122: 2239-2250.
- Cao, Y., Chen, H., Zhou, L., Chiang, M. K., Anand-Apte, B., Weatherbee, J. A., Wang, Y., Fang, F., Flanagan, J. G., and Tsang, M. L. (1996) Heterodimers of placenta growth factor/ vascular endothelial growth factor. Endothelial activity, tumor cell expression, and high affinity binding to Flk-1/KDR. J. Biol. Chem. 271: 3154-3162.
- Chiang, M. K. and Flanagan, J. G. (1995) Interactions between the Flk-1 receptor, vascular endothelial growth factor, and cell surface proteoglycan identified with a soluble receptor reagent. Growth Factors 12: 1-10.
- Skoda, R. C., Seldin, D. C., Chiang, M. K., Peichel, C. L., Vogt, T. F., and Leder, P. (1993) Murine c-mpl: a member of the hematopoietic growth factor receptor superfamily that transduces a proliferative signal. EMBO 12: 2645- 2653.
